Selection of short how-to clips related to XRD Nanopowder Processor
- Sample DataStore 9.2 – result Extraction + XRD
Video about extraction of search results and pasting into Excel - XRD NanoProcessor -1- Introduction
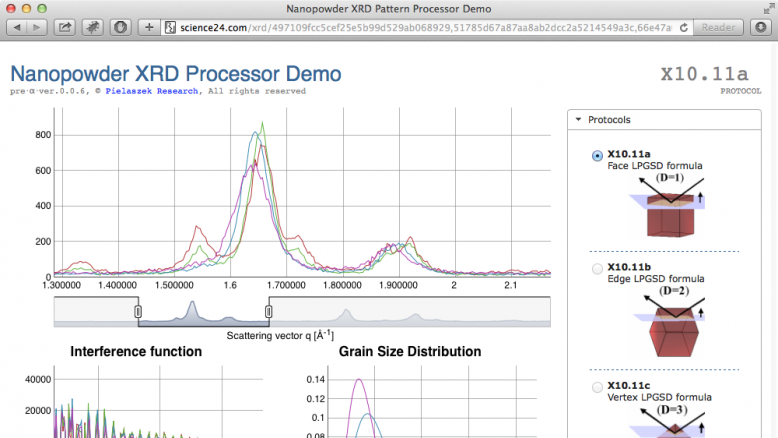
Video introduction to X-Ray Diffraction (XRD) Nano Processor Demo. This web application allows for Drag & Drop processing of nanopowder (nanocrystalline) diffraction data and obtain grain size distribution in few seconds. - XRD NanoProcessor -2- Theory
Video introduction to X-Ray Diffraction (XRD) Nano Processor Demo. This web application allows for Drag & Drop processing of nanopowder (nanocrystalline) diffraction data and obtain grain size distribution in few seconds. Commentary on diffraction theory used for processing the XRD data. - XRD NanoProcessor -3- Assumptions
Video introduction to X-Ray Diffraction (XRD) Nano Processor Demo. This web application allows for Drag & Drop processing of nanopowder (nanocrystalline) diffraction data and obtain grain size distribution in few seconds. Commentary on assumptions and limitations while processing the XRD data. - XRD NanoProcessor -4- Band and Coherence distance
Video introduction to X-Ray Diffraction (XRD) Nano Processor Demo. This web application allows for Drag & Drop processing of nanopowder (nanocrystalline) diffraction data and obtain grain size distribution in few seconds. Commentary on band and coherence distance parameters of the XRD data. - XRD NanoProcessor -5- Sharing results
Video introduction to X-Ray Diffraction (XRD) Nano Processor Demo. This web application allows for Drag & Drop processing of nanopowder (nanocrystalline) diffraction data and obtain grain size distribution in few seconds. Commentary on sharing results of processing of the XRD data. - XRD NanoProcessor -6- Examples
Few example of data processing with X-Ray Diffraction (XRD) Nano Processor Demo. Good data and success story is shown in this clip. - XRD NanoProcessor -7- Bad Data Examples
Example of processing bad data with X-Ray Diffraction (XRD) Nano Processor Demo. The routine should break calculation to avoid seriously incorrect conclusions.

Hello,
I find your method to estimate GSD very interesting. I gave read the related papers by R. Pielaszek et al. and try to look at the Pielaszek’s dissertation (unfortunately in Polish) , still I have some questions about the application of and the models and protocols used in the “Nanopowder XRD Processor Demo”:
1. I do not see any parameters in the analytical line profile LP(q,, sigma) or GSD(R;,sigma) to account for the instrumental broadening (B_ins.), so how one can take B_ins into account to get accurate and sigma?
2. To fit LP(q,, sigma) to an XRD peak one need to approximate and sigma, or at leat write them as function of the peak parameters (height, FWHM, position, area …). Which relation did you use to approximate initial value of ?
4. How to choose the peaks to fit? and It is possible to fit all peaks for a phase and draw GSD(R)??
3. The “Nanopowder XRD Processor Demo” site do not provide any informations on the models implemented in your protocols and the difference between them. This is hindering the use and citation of that tool.
A last comment I hope that you accept it take it seriously. The “black box” strategy in developing models, algorithms and programs for academic purposes is not a good idea, especially at the beginning. And I think that is why many nice tool such yours do not get the required attention by scientific community. Any student or researcher should know the finest details about the tools he use, the lack of such details push them to use other tools or reinvent the wheel again.
I hope that we can see a decent open-source version of your tool soon.
Best regards.
Many thanks for comments. Answering your questions:
True. Present version (July 2016) does not account for instrumental broadening. As you can see in “Experimental” panel, there’s a subform (now disabled) to drop a reference pattern. We look for funding (e.g. some EU project) to implement this and processing of amorphous phase, which now inhibits many patters from being accepted. Normally, we join some projects providing scientific databases (http://pielaszek.com/about-us/services/sample-management/, http://pielaszek.com/sample-datastore-9-2-video-tutorials/) and try to develop XRD tools as well.
None. That’s the point. There is no peak fit at all. Too unstable to work in a fully automated toll. Instead, we do a math trick similar to “hybridisation” in quantum chemistry. No physical meaning whatsoever, just abstract math, but handy. We merge all peaks into one and process this “super peak” alone. That’s why in presence of two (or more) phases this protocol stops processing (amorphous is also considered “a phase”). On the other hand it seems to work ok for convoluted stuff, see Fig.1.8 and Fig.1.16 in attached book.
Today the only way os to drop part of pattern (cut a peak). Full (and automatic) phase recognition is doable, but… (see funding remarks above).
Citing two (old) papers referenced on the front page and the web address (science24.com/xrd) itself for us is ok. Beside LPGSD formulas there is no specific model to cite. Well, maybe this hybridisation. But that would take a book to describe with reproducible details. Protocol names Xnn.nn enumerate computing code, so one is sure that the same protocol will always produce the same results (including bugs/errors). That’s for industry.
The tool isn’t meant for academia. However, mostly academia uses that :). Idea is not to require any user input (besides the pattern). Imagine factory of nanomaterials with attached XRD for quality control at the end pipe. Underlying engine must return valid data without asking a question.
On the other hand, there is nothing to hide. Two new books undrego: Nanoparticles (Nanoczastki) from Polish Scientific Publishers (PWN) and other from Warsaw University Press. In the first I gave short (~30 pages) resume since it’s a “brick” from ~15 authors. The second is a bit more detailed, you got that attached (working copy). Both in Polish, unfortunately, since these were the only publishing offers I got so far.
Publishing papers is worthless – theoretical physics is too demanding in terms of space. Besides, I hate giving work they immediately hide (or pay $40 to read few pages).
Instead, I’ll be more than happy to answer any specific question in this blog. Valid questions with answers will go to public with no delay, free of charge.
Existing tool is a server-side code, a mixture of many packages and languages. No wrapable as a standalone project. Also, for licensing reasons (not all components belong to us). However, we’ll provide essential knowledge for external developers without reluctance.
Cheers,
roman